Photoactivated Localization Microscopy, or PALM imaging, is a super-resolution technique that relies on the random activation of fluorophores to spatially resolve molecular details (Betzig et al. 2006). In brief, upon activation with the adequate laser, a sparse number of fluorophores emit for a short period before bleaching. Because fluorophores are only activated at the same time in small numbers until all have emitted, it is possible to localize and track single molecules over time.
Similarly to any single molecule localization microscopy (SMLM) techniques, the key in PALM imaging is to use the right fluorophores. PALM microscopy is particularly recommended for live-cell imaging, counting and tracking of molecules in cells or in solution, as most fluorophores are normally only activated once – except those with photoswitchable properties (see below). On the contrary, dSTORM microscopy, where phototoxic buffers and high-power lasers are applied to induce fluorescent dyes to go through several on and off states before bleaching, is mostly suitable for fixed samples. In this post, we summarize some of the different photoactivatable, photoconvertible or photoswitchable fluorophores that can be used for PALM imaging in live cells, considering your experimental requirements and protein cellular localization.
Photoactivatable (PA) fluorophores
These emit light upon activation, most commonly with UV irradiation, and are the most commonly used fluorophores in PALM imaging. They are photoactivatable versions of commonly used fluorescent proteins (FPs), which can be fused to the protein of interest. In general, it is important to keep in mind that photoactivatable fluorophores are dark by nature and imaging can be tricky at first (the cells will be dark too!), requiring some optimization.
- PA-GFP: fluorophore excitation 504nm, emission 517nm (green), photoactivated with UV-Violet (405nm) for high emission or blue (488nm) laser for low emission. It is recommended for certain live cell and multicolor experiments (Patterson and Lippincott-Schwartz 2002). Offers less flexibility and complexity in experimental design than others. It can dimerize, some point mutants have been designed to prevent this./li>
- PA-TagRFP: fluorophore excitation 562nm, emission 595nm (red), photoactivated with UV-Violet (405nm). Very bright, photostable and of monomeric nature, excellent protein tag for both conventional microscopy and super-resolution PALM microscopy (Subach et al. 2010). Good for two-color PALM imaging.
- PA-mCherry1: fluorophore excitation 570nm, emission 596nm (red), photoactivated with UV-Violet (405nm). Rapidly-maturing monomer with high acid sensitivity (Subach et al. 2009). Good for two-color PALM imaging. However, its low photon count makes it one of the worst fluorophores for PALM tracking.
- PA-mKate2: fluorophore excitation 586nm, emission 628nm (far-red), different PA mechanisms, can be photoactivated with UV-Violet (405nm). Monomeric far-red, higher pH stability and slower photobleaching than PA-mCherry1 (Gunewardene et al. 2011).
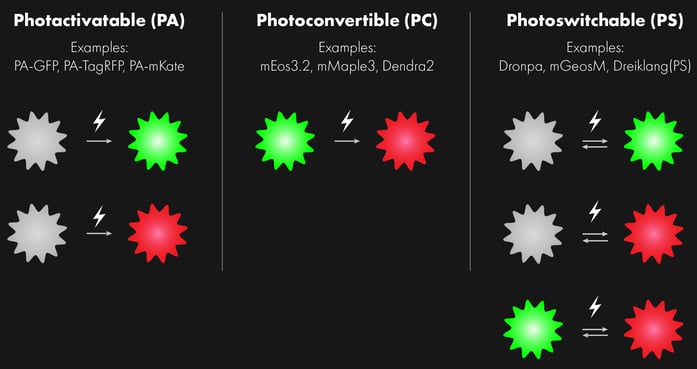

Photoconvertible (PC) fluorophores
These are fluorescent proteins that already emit fluorescence in their non-converted state and that change their emission spectrum upon activation, most commonly with UV light. The conversion is irreversible. They offer accurate localization thanks to high photon outputs. Some of the widely used PC fluorophores, most from green to red, are:
- PS-CFP2: irreversible photo-switching from cyan to green, excitation 400nm (cyan) and 490nm (green), emission 468nm (cyan) and 511nm (green), photoconversion with UV-Violet (405nm). Easily imaged without photo-activation, high pH stability and good candidate for multicolor imaging.
- Kaede: excitation 508nm (green) and 572nm (red), emission 518nm (green) and 582nm (red), photoconversion with UV-Violet (405nm). First PC-fluorophore. Its tetrameric nature makes it challenging to use for live-cell imaging experiments, as oligomerization can cause artefacts and image misinterpretation.
- mEos2: excitation 506nm (green) and 573nm (red), emission 519nm (green) and 584nm (red), photoconversion with UV-Violet (405nm). Better than its earlier version tdEOS (which tetramerizes) but tends to form dimers and high order oligomers at high concentrations, which makes it unsuitable for labeling membrane proteins. Recommended to use later version mEos3.2 or mMaple3 instead (Wang et al. 2014).
- mEos3.2: excitation 507nm (green) and 572nm (red), emission 516nm (green) and 580nm (red), photoconversion with UV-Violet (405nm). mEos3.2 is truly monomeric, brighter, matures faster and exhibits higher photon budget and label density (Zhang et al. 2012). Along with mMaple3, considered best of the green to red PC-FPs.
- mEos4b: excitation 505nm (green) and 569nm (red), emission 516nm (green) and 581nm (red), photoconversion with UV-Violet (405nm). Osmium tetroxide resistant, engineered for correlative EM imaging.
- mMaple3: excitation 489nm (green) and 566nm (red), emission 505nm (green) and 583nm (red), photoconversion with UV-Violet (405nm). Most recently introduced photoconvertible FP for single-molecule localization, very high detectable FP vs total FP ratio (McEvoy et al. 2012).
- Dendra2: excitation 490nm (green) and 553nm (red), emission 507nm (green) and 573nm (red), photoactivated with UV-Violet (405nm) or blue (488nm) laser. Blue light activation and good performance makes Dendra2 a good candidate for live-cell single-molecule localization microscopy. Recent efforts have focused on improving this fluorophore by using primed photoconversion (PC) in combination with UV-photoactivation (Virant et al. 2017), so that fluorescent proteins can be specifically photoconverted by different light wavelengths using PC and UV-activation modes but emit fluorescence in the same spectral emission channel. Another approach has proven to reduce phototoxicity in live‐cell single‐molecule imaging by introducing point mutations to facilitate photoconversion of Dendra2 using blue and complementary near‐infrared light (Turkowyd et al. 2017).
- PSmOrange: excitation 561nm (orange) and 636nm (far-red), emission 565nm (orange) and 662nm (far-red), photoconversion with UV-Violet (405nm) or blue (488nm laser). PSmOrange photoswitching occurs via a two-step photo-oxidation process, which causes cleavage of the polypeptide backbone. The red-shifted spectra of both forms of PSmOrange enable its simultaneous use with cyan-to-green photoswitchable proteins, making it good for multi-color super-resolution microscopy to study different markers (Subach et al. 2011).
Photoswitchable (PS) fluorophores
Contrary to photoactivatable fluorophores, photoswitchable fluorophores are reversible and able to shuttle between a non-fluorescent and a fluorescent state, cycles can be repeated several hundred times without photo-bleaching. They tend to be bright and, therefore, give high localization precision. However, if you are aiming at study the dynamics of individually labeled proteins over time in live PALM imaging, it will be better to use photoactivatable or photoconvertible fluorescent proteins, which will only emit once.
- Dronpa: excitation 503nm, emission 518nm (green), photoactivated with UV-Violet (405nm). Reversibly switchable from dark to green. We would recommend using PS-CFP2 instead for most single-molecule localization applications. Extensive mutagenesis of Dronpa has been used to generate additional fluorophores variants with faster photoswitching kinetics, such as rsFastlime and Padron, which displays a switching behavior that is reversed to that of all green fluorescent reversibly switchable fluorescent proteins known to date (Andresen et al. 2008).
- mGeosM: excitation 503nm, emission 514nm (green), photoactivated with UV-Violet (405nm). Better than Dronpa, but with a higher duty cycle than PS-CFP2. mGeos was created by mutating the first amino acids in the chromophore of the photoconvertable fluorescence protein mEos2 (McKinney et al. 2009).
- Dreiklang: excitation 515nm, emission 529nm (green), photo-deactivated with UV-Violet (405nm). Reversible on-off switching in living cells is accomplished at two different wavelengths of 365nm and 405nm, respectively, whereas fluorescence is elicited at 515 nm (Brakemann et al. 2011). It has high acid sensitivity, low contrast at low pH values, so it will not work in all cell compartments.
- mIrisGFP: excitation 486nm (green) and 516nm (red), emission 546nm (green) and 578nm (red). Photoconversion with UV-Violet (405nm) from dark to green and green to red; or photoconversion with blue (488 nm) for dark to red. Photoconvertable from green to red state, both green and red states reversibly photoswitchable to dark state. Demonstrated for combined SMLM with pulse-chase experiments.
- NijiGFP: excitation 469nm (green) and 526nm (red), emission 507nm (green) and 569nm (red). Photoconversion with UV-Violet (405nm) from dark to green and green to red; or photoconversion with blue (488 nm) for dark to red. Photoconvertable from green to red state, both green and red states are reversibly photoswitchable to a dark state (Zhou and Lin 2013).
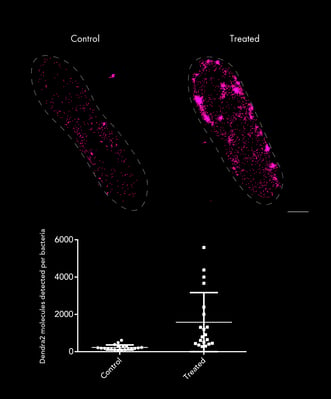
PALM imaging of Escherichia coli bacteria control (left) or treated with antibiotic (right), in which the number of molecules of Transcription Factor-Dendra2 fusion proteins were quantified for each condition. Sample provided by Prof. Michael Schlierf's lab at the B CUBE - Center for Molecular Bioengineering TU Dresden.
Alternative labeling methods: Halo tags
An alternative labeling approach that has gained adepts in recent years is the use of hybrid systems; a combination of genetically-encoded protein tags with a separate cell-permeable synthetic fluorescent ligand. These include proteins tagged with SNAP tag, CLIP tag and the most used HaloTag® – a system that uses a range of fluorescence ligands, such as tetramethylrhodamine (TMR) and Janelia Fluor® 549 or 646 (Grimm et al. 2016). These tags combine the advantages of genetically encoded fluorescent proteins with the superior brightness of synthetic dyes and photostability (less prone to photobleaching than FPs), but require a laborious labeling process (Los et al. 2008). If optimized, these systems allow researchers to carry out high-speed and dual-color single-molecule tracking in live cells (Banaz et al. 2018).
It is not only genetically-encoded fluorescent proteins can be used for PALM imaging. Synthetic dyes or quantum dots conjugated to antibodies can also be used. However, these are only suitable for fixed samples, not appropriate for live-cell imaging and specific targeting of the protein of interest can be more challenging. For their use, cells have to be permeabilized to ensure access through the plasma membrane and into the cell interior, and fluorescence intensity levels can vary depending on the concentration of labeled secondary antibody or fluorescent dye. 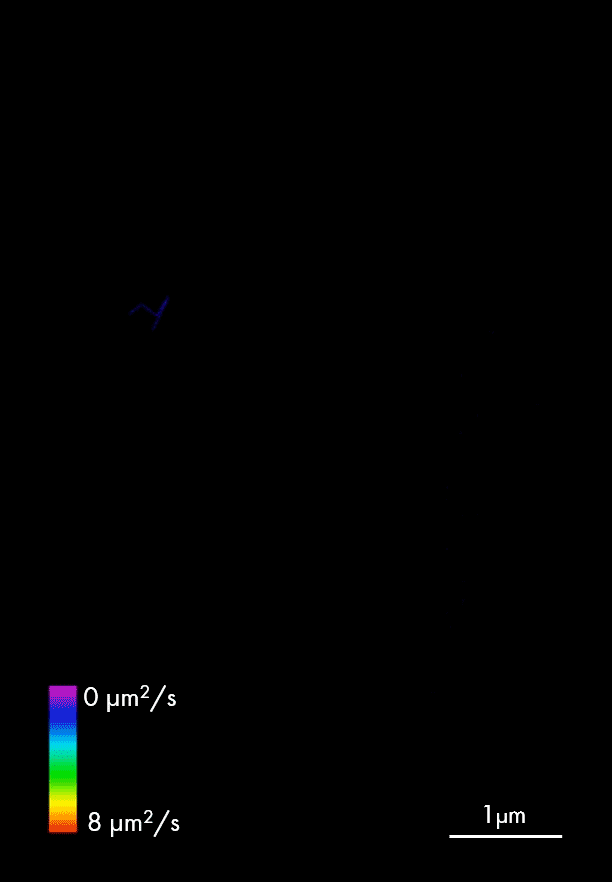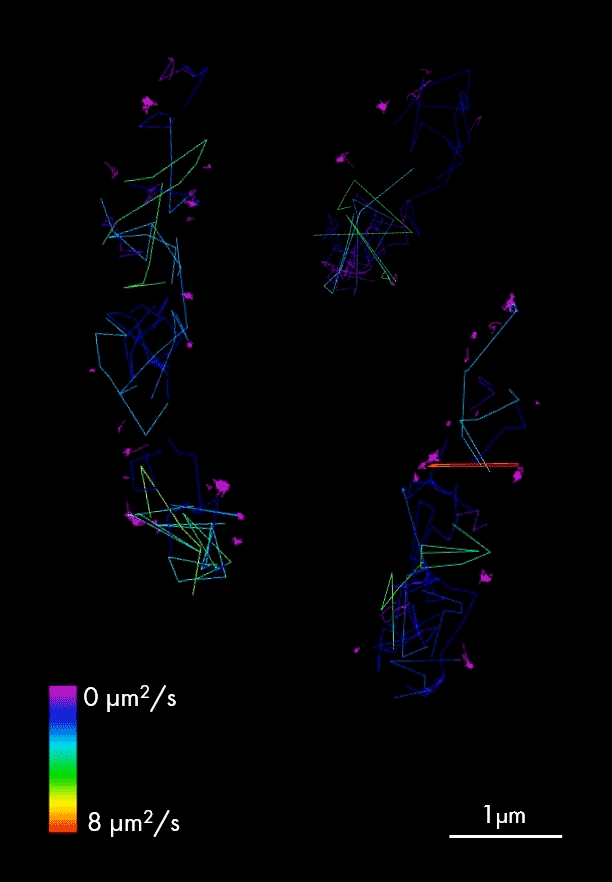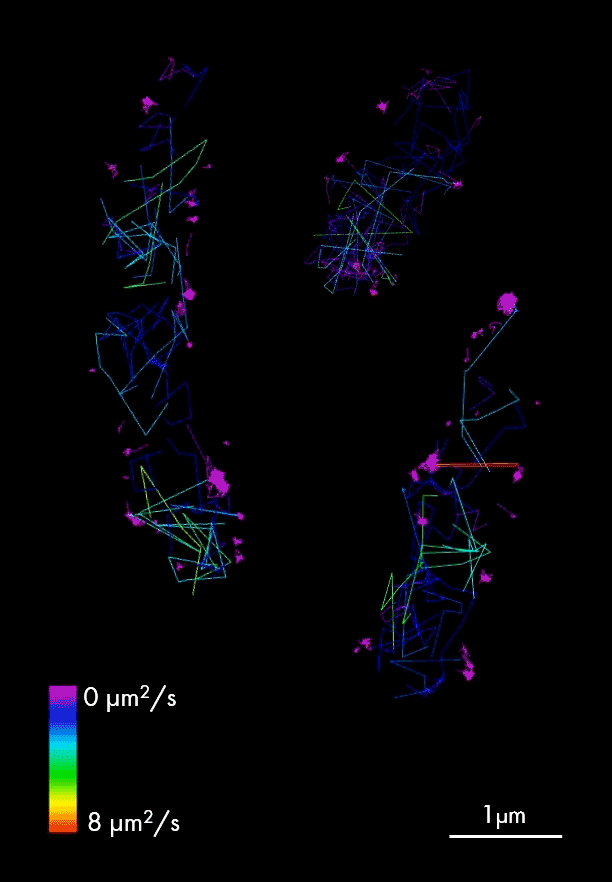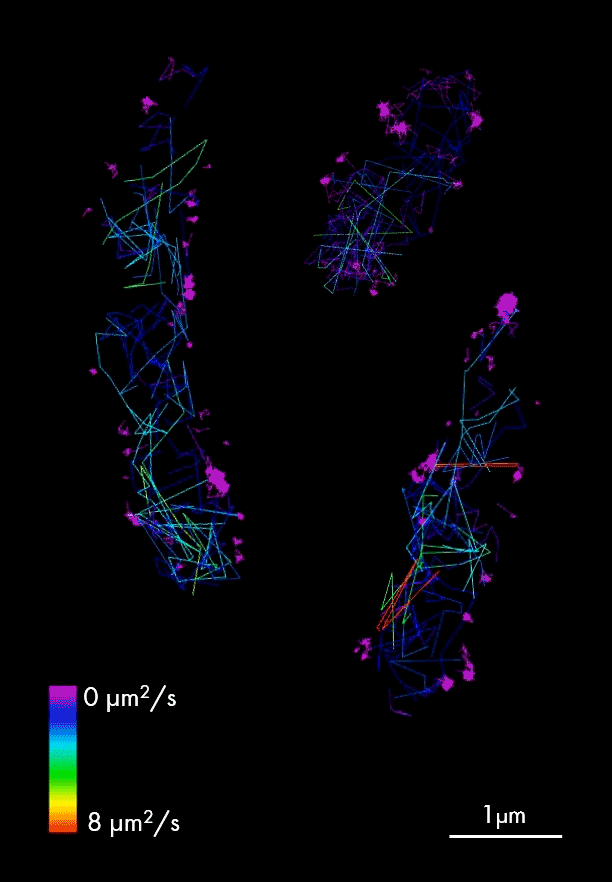
Single particle tracking of T3SS-HALO stained with JF-549 in live bacteria. Sample provided by Dr. A. Diepold, Max for Terrestrial Microbiology, Marburg (Germany). Scale bar is 1µm.
For further literature insights, read a recent comprehensive review on photosensitive fluorophores for single-molecule localization microscopy (SMLM). You can also read more about the basics of PALM microscopy or single-particle tracking.
Are you new to super-resolution microscopy? Unsure of what is the best techniques for you? Download our techniques table to help you choose between the Nanoimager's key super-resolution microscopy techniques. It covers core techniques including dSTORM, PALM imaging, Single-Particle Tracking, SIM and smFRET.
You can also download a summary guide to the most popular fluorophores for PALM imaging based on this blog post. Happy imaging!
Share this article:
