Single-Molecule Localization Microscopy (SMLM) techniques such as Direct Stochastic Optical Reconstruction Microscopy (dSTORM) and DNA-based point accumulation for imaging in nanoscale topography (DNA-PAINT) are powerful tools to investigate structures at the 20 nm resolution scale. This resolution gain compared to diffraction-limited techniques (resolution approximately 250 nm) comes at the expense of a high sensitivity for background fluorescence and, sometimes, imaging artifacts. Designing the right controls oftentimes aids in the optimization of sample preparation protocols for SMLM.1
Considerations in the design of your first super-resolution experiment
Understanding what to expect from SMLM is the first step to acquire nanoscale resolution images. To get familiar with the technique, especially the fluorophore choice, sample labeling, and the correct buffer conditions (blinking buffers for dSTORM or optimal binding buffer for DNA-PAINT), we recommend testing your chosen dye by labeling a known structure with a distinctive morphology. This can be, for example, staining mitochondria or tubulin structures in cells, which have an easily recognizable structure.
For a selection of the most popular fluorophores for dSTORM and the recommended buffers, download our Popular fluorophores for dSTORM imaging overview. In contrast to dSTORM, PAINT imaging requires bright and photostable dyes 2. Furthermore, the oligonucleotide sequence and imaging buffer can be optimized to achieve optimal imaging speed and resolution.3
Sample labeling is crucial to optimally reconstruct your desired structure. This can be done by optimizing the antibody concentration for labeling. A high labeling concentration ensures that all the available epitopes are labeled. For reference, follow the dSTORM sample preparation workflow.
Using a well-known standard will also provide an idea about the sample blinking behavior and the resolution that can be obtained. In dSTORM specifically, it is important that the buffer is optimal for your chosen dye to obtain the best results. ONI’s BCubed buffer efficiently regulates blinking behavior and helps achieve high-quality super-resolution images.
Essential controls for one-color SMLM imaging
The most common labeling strategy for SMLM is immunolabeling (or immunofluorescence staining), involving the staining of your target of interest with either a primary antibody (direct immunolabeling) or a combination of primary and secondary antibodies (indirect immunolabeling). For example, immunolabeling can be applied in the quantification of cell surface biomarkers or the characterization of extracellular vesicles (EVs).
It is important to run the controls using the exact same imaging parameters as the sample (frames, exposure time and illumination angle), to keep everything consistent. We can distinguish between positive and negative controls for 1 color SMLM using immunolabeling:
Positive controls for SMLM
When imaging an unknown sample it is generally helpful to include a positive control. This usually consists of a sample where the target is expressed in high quantities, for example, an overexpressed cell receptor. A good positive control can also be a well-characterized sample standard. In case of EV imaging, ONI’s EV Profiler kit provides an EV standard positive control. If there is a lack of localizations in the positive control, it might indicate that a step in the sample preparation went wrong, such as the surface preparation, sample attachment to the coverslip or the sample labeling protocol.
Negative controls for SMLM
- Unlabeled structure. Adding a control condition that includes your sample without any fluorophore aids in identifying background fluorescence from the sample itself or the imaging buffer. This can be either your sample alone without staining or adding your primary (and secondary if applicable) antibody without fluorophores. This negative control is critical, especially in cell imaging, as some cell types or cell-secreted vesicles can show autofluorescence mainly in the 488 or 561 channel.
- Isotype control. Addition of antibodies that have no interaction with any known targets within your sample. Notably, the isotype control antibody should be of the same species and concentration as your specific labeling antibody. This type of negative control helps to evaluate the level of non-specific background antibody staining. Ideally, only sparse individual localizations should appear in the isotype control sample.
- Unspecific binding of secondary antibodies (in case of indirect immunolabeling). Using a primary antibody in combination with a secondary antibody to label the target of interest is a common approach to amplify the fluorescent signal. This is because multiple fluorescently labeled secondary antibodies can bind to a single primary antibody. However, this might also lead to a significant increase in background fluorescence. In this scenario it is important to include a control of the secondary antibody only (without primary antibody) to evaluate the unspecific binding to the sample. It is worth noting that secondary antibodies can also form aggregates that could be identified as false clusters in your SMLM image.
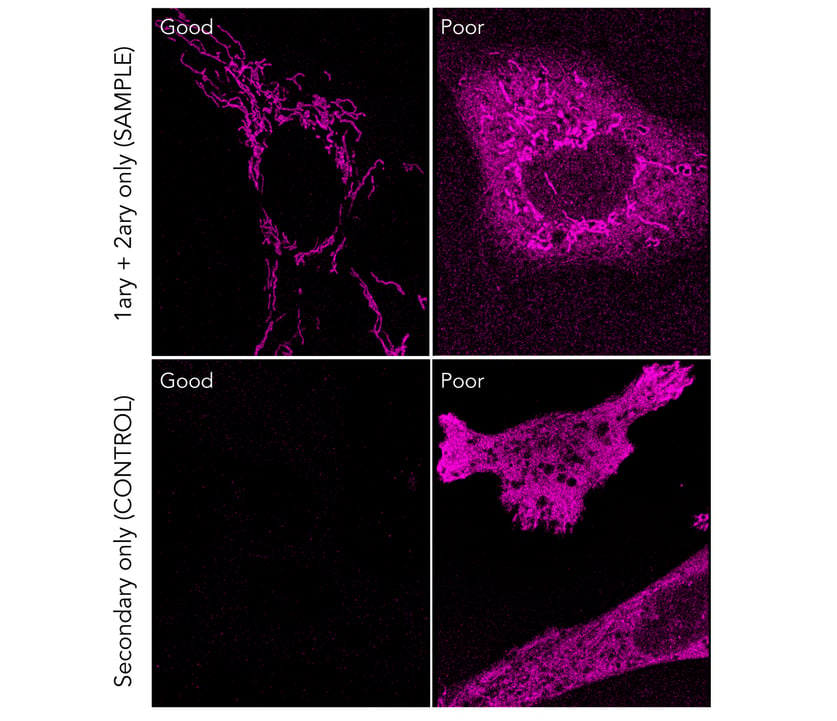
Figure 1. Top panel: dSTORM image of cells stained with an anti-TOMM20 antibody, followed by a secondary antibody conjugated to AlexaFluor-647. An example of a staining with minimal background can be found on the left panel, where mitochondria are nicely visualized. Insufficient blocking can lead to nonspecific binding of the secondary probes, increasing the background level and ultimately lead to no distinction between the specific and nonspecific signal (right panel). Bottom panel: Background fluorescence signal of control cells stained only with the secondary antibody conjugated to AlexaFluor-647 (in the absence of a target-specific primary antibody). An example of a good negative control can be seen in the left panel, while insufficient blocking shows nonspecific binding of the secondary probes, leading to false positive signal (right panel). All dSTORM images were captured during 10000 frames at a 30 ms exposure and 55 mW laser power using ONI’s Nanoimager. Contrast settings were set the same for all conditions.
Here are our two main tips for optimizing the labeling density of your target of interest:
- Perform a titration of the primary antibody to achieve the best labeling density for low- or high-expression targets. Use 5 µg/mL of primary antibody as a starting point.
- As a rule of thumb, for STORM imaging, you should likely be using at least twice the concentration of antibody that you use in conventional immunofluorescence microscopy (confocal or widefield).
Negative controls for DNA-PAINT
DNA-PAINT uses transient binding of complementary DNA strands to achieve the blinking behavior needed for SMLM. The target of interest is usually labeled with an antibody coupled to a single DNA docking strand while its complementary imager strand is coupled to a dye freely floating in solution. Non-specific binding events can originate from the unspecific attachment of the imager strand to the sample. Thus, two negative control conditions are essential to include in DNA-PAINT experiments:
- Wrong pairing. This condition includes the addition of an imager strand that has no base pair complementarity to the docking strand of the labeled structure of interest. This control determines the amount of nonspecific background localizations and validates that the chosen docking-imaging strand interaction is specific.
- Imager only. The addition of the imager strand to a sample that contains no docking strands can give an insight into the nonspecific binding of the imager strand to the sample. In some cases, it might be necessary to add a blocking step in your sample preparation protocol to ensure that the binding of the imager alone to the sample is minimal. Furthermore, the imager concentration should be optimized to avoid a high background fluorescence in solution, that can affect the signal-to-noise ratio and the localization precision of specific binding events. While background binding and fluorescence can be reduced by reducing the imager concentration in solution, this comes with a longer imaging acquisition time tradeoff.
Advanced imaging controls in SMLM
Additional controls need to be considered for specific SMLM experiments, including multicolor SMLM imaging and the evaluation of unspecific free dye attachment to the sample.
-
Controls for multicolor SMLM. Choosing your dye combinations carefully to avoid cross-talk between different channels. We recommend when designing your multicolor experiment to add single label controls to understand the number of localizations present in the non-labeled channels. Generally, a low level of bleed-through between channels can be corrected for by using the right post-processing settings, such as a photon count threshold. This ensures that only bright localizations are detected and counted which correspond to the labeled channel.
In the case of multiplexing in DNA-PAINT it is worth investigating the specificity of the DNA sequences to confirm that the different sequences used do not interact with each other. Similarly to the wrong pairing control, it is recommended to test that imager sequence 1 does not bind specifically to the docking sequence 2 and that the imager 2 does not bind to docking 1 in a two-color DNA-PAINT example.
-
Unspecific dye attachment. Some dyes, normally very hydrophobic in nature, are known to be sticky towards biological membranes such as vesicles or cell membranes. In case of doubt, when using a non-standard dye or an unknown biological sample, it might be recommended to perform a control condition where only the dye molecule is added to the sample so that the level of non-specific dye binding can be determined.4
Concluding remarks
Background localizations are a part of every super-resolution microscopist's data, due to the sensitivity of SMLM technique. Using the appropriate controls, scientists can make sure that what they are seeing is a target-specific signal and validate the sample preparation protocols. However, avoiding all background localizations is a hard and nearly impossible aspiration. For quantitative purposes, scientists have developed strategies to minimize the background counting, such as filtering and clustering. ONI’s novel analysis software CODI is designed to make visualization and analysis of SMLM data easy, accessible and collaborative.
For getting started into SMLS imaging, also check out our recent Dos and Don’ts for mastering dSTORM imaging article and take a look at The Ins and Outs of mastering dSTORM imaging webinar.
If you are eager to learn more, do not hesitate to visit our website and get in touch with us at hi@team.oni.
References
- Jimenez A, Friedl K, Leterrier C. About samples, giving examples: Optimized Single Molecule Localization Microscopy. Methods. 2020; 174: 100-114. https://doi.org/10.1016/j.ymeth.2019.05.008.
- Tholen, MME, Tas, RP, Wang, Y, Albertazzi, L. Beyond DNA: probes for PAINT super-resolution microscopy. Chem. Commun. 2023, 59, 8332-8342. https://doi.org/10.1039/D3CC00757J
- Schueder F, Stein J, Stehr F et al. An order of magnitude faster DNA-PAINT imaging by optimized sequence design and buffer conditions. Nat Methods. 2019; 16: 1101–1104. https://doi.org/10.1038/s41592-019-0584-7
- Hughes LD, Rawle RJ, Boxer SG. Choose your label wisely: water-soluble fluorophores often interact with lipid bilayers. PLOS one. 2014; 9(2): e87649. https://doi.org/10.1371/journal.pone.0087649
Share this article: