Challenge: Understand viral particle structure and localization
 Viral particles vary greatly in size, ranging from approximately 20 to 300 nm in diameter, which can be below or close to the resolution limit of conventional fluorescence microscopy. Consequently, there are many mechanistic and functional characteristics of viral particles that are yet to be elucidated.
Viral particles vary greatly in size, ranging from approximately 20 to 300 nm in diameter, which can be below or close to the resolution limit of conventional fluorescence microscopy. Consequently, there are many mechanistic and functional characteristics of viral particles that are yet to be elucidated.
Recently, super-resolution imaging techniques have been employed to study viral particles and their contents at single-molecule level.
Solution with the Nanoimager: Imaging viral particles using dSTORM microscopy
Super-resolution microscopy opened a plethora of possibilities to understand the molecular mechanisms underlying viral particle behavior in solution and in cells. Using dSTORM we can now gain detailed knowledge of how and where individual viruses are arranged, localized or can form aggregates, for instance.
Information that supports this could include the organization of molecules on membranes or inside viral particles, their interactions, structural preservation and their local accumulations. Such analysis can be run on a coverslip or in a biological context. Researchers can analyze these structural changes arising in response to a stimulus, a change in the cell environment or a particular cell cycle stage.
The Nanoimager, through its dSTORM microscopy capabilities, offers the opportunity to see inside the cell or look at purified viral particles with a resolution of up to 20 nm using localization-based super-resolution microscopy.
With immunofluorescence labelling, the spatial relationship of up to four molecular species can be imaged with the four laser lines. Additionally, the Nanoimager not only presents super-resolved images of cellular structures, but it offers several tools for quantifying this information.
With its extreme sensitivity and easy to use analysis tools, the Nanoimager can quantitatively discriminate between positive and negative controls, and perform single-particle tracking. Additionally, it offers the advantage of a large field of view and simultaneous dual channel imaging, which, together with real-time rendering, can drastically reduce the acquisition time and interpretation of the obtained results.
Case Study: Analysing viral particle architecture with dSTORM microscopy
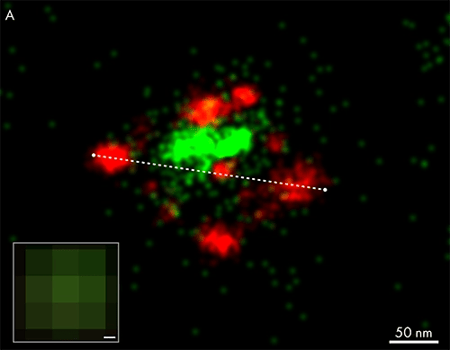 The benefit of super-resolution imaging with dSTORM microscopy is presented in the figure to the right, where the viral surface glycoprotein (red) and cellular host factors incorporated into the particle (green), of a single-virus particle adhered to coverslip, were investigated by labelling with antibodies conjugated to AF647(red) and AF488 (green). The histogram across the viral particle (panel B) shows its width to be around 190 nm, and the cellular host factor accumulating inside the particle. This level of information is not obtainable in a conventional fluorescence widefield image (panel A, inset).
The benefit of super-resolution imaging with dSTORM microscopy is presented in the figure to the right, where the viral surface glycoprotein (red) and cellular host factors incorporated into the particle (green), of a single-virus particle adhered to coverslip, were investigated by labelling with antibodies conjugated to AF647(red) and AF488 (green). The histogram across the viral particle (panel B) shows its width to be around 190 nm, and the cellular host factor accumulating inside the particle. This level of information is not obtainable in a conventional fluorescence widefield image (panel A, inset).
Learn more about the features of the Nanoimager’s NimOS software and the super-resolution microscopy techniques that our microscope supports.
Share this article: