Challenge: Understanding protein complexes
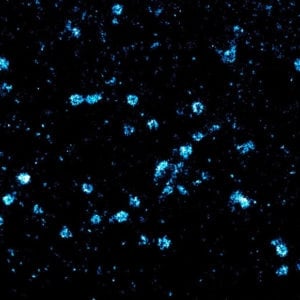 Small protein complexes are ubiquitous across molecular biology, they are involved in transport and translocation of molecules across membranes, signaling and various important mechanisms. Such processes are vital for the proper functioning of cells and have disastrous consequences when they go wrong, from heart disease to mental illnesses. Understanding the structure, interacting components and location of these nanometer-sized complexes is a difficult task.
Small protein complexes are ubiquitous across molecular biology, they are involved in transport and translocation of molecules across membranes, signaling and various important mechanisms. Such processes are vital for the proper functioning of cells and have disastrous consequences when they go wrong, from heart disease to mental illnesses. Understanding the structure, interacting components and location of these nanometer-sized complexes is a difficult task.
Solution with the Nanoimager: Starting protein complexes at the nanoscale level
The Nanoimager boasts several features to support the study of protein complexes at the nanoscale level:
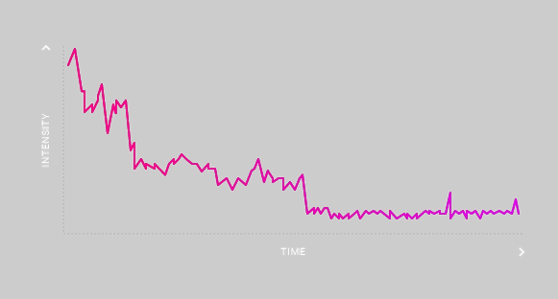
- Counting the number of subunits in a complex by intensity – the number of labeled proteins is equal to the sum of the intensity divided by that of a single fluorophore.
- Measuring the dynamics of assembling complexes, using single-particle tracking and/or smFRET.
- Counting subunits by the number of decremental steps as labeled proteins in a complex go dark due to photobleaching, as shown in the figure.
- Investigating the structure of protein complexes by super-resolution imaging, as has been rigorously demonstrated in the past with the nuclear pore complex and larger structures such as the cytoskeleton (see the clathrin pits image above).
Famous examples of these methods applied to protein complexes include counting the subunits in the bacterial flagellar motor and in ion channels, as well as measuring the aggregation of amyloid beta associated with Alzheimer’s.
Case Study: Resolving protein complex assembly with the Nanoimager
 In this example, the Nanoimager was used to study the assembly of the Twin-arginine translocation (Tat) complex. The Tat complex is notable for its capacity to transport folded proteins across the cytoplasmic membrane in bacteria; as it is not present in mammalian cells, it is a potential target for anti-microbial drugs.
In this example, the Nanoimager was used to study the assembly of the Twin-arginine translocation (Tat) complex. The Tat complex is notable for its capacity to transport folded proteins across the cytoplasmic membrane in bacteria; as it is not present in mammalian cells, it is a potential target for anti-microbial drugs.
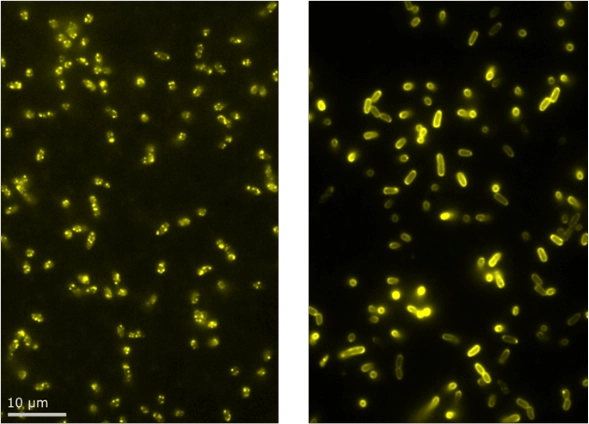
The figure shows E. coli cells expressing a YFP-labeled component of the Tat complex. Each Tat complex contains multiple copies of this component, so when the complexes assemble they appear as distinct bright spots; each spot in the image contains multiple YFPs.
A mutation in the Tat complex prevents it from assembling, therefore in the second phenotype (shown on the right), the YFP are diffusing single molecules which appear as a more homogeneous distribution on the membrane under particular exposure settings. In the 2D projection shown here, this homogeneous distribution gives the effect of a halo around the bacteria. Ultimately, the assembled complexes are distinguished by their intensity and distribution and the mutation is shown to prevent normal assembly of the complex.
Learn more about the Nanoimager and the super-resolution techniques that it supports.
Figure (right): E. coli cells expressing a YFP-labeled component of the Tat complex
Share this article: